What ESMValTool can do for you#
The ESMValTool applies a great variety of standard diagnostics and metrics, and produces a collection of netCDF and graphical files (plots). Thus, the tool needs a certain amount of input from the user so that it can:
establish the correct input and output parameters and the structured workflow;
acquire the correct data;
execute the workflow; and
output the desired collective data and media.
To facilitate these four steps, the user has control over the tool via the configuration and the recipe. The configuration sets user and site-specific parameters (like input and output paths, desired output graphical formats, logging level, etc.), whereas the recipe file sets data, preprocessing and diagnostic-specific parameters (data parameters grouped in the datasets sections, preprocessing steps for various preprocessors sections, variables’ parameters and diagnostic-specific instructions grouped in the diagnostics sections). The configuration file may be used for a very large number of runs with very minimal changes since most of the parameters it sets are recyclable; the recipe file can be used for a large number of applications, since it may include as many datasets, preprocessors and diagnostics sections as the user deems useful.
Once the configuration files and the recipe are at hand, the user can start the tool. A schematic overview of the ESMValTool workflow is depicted in the figure below.
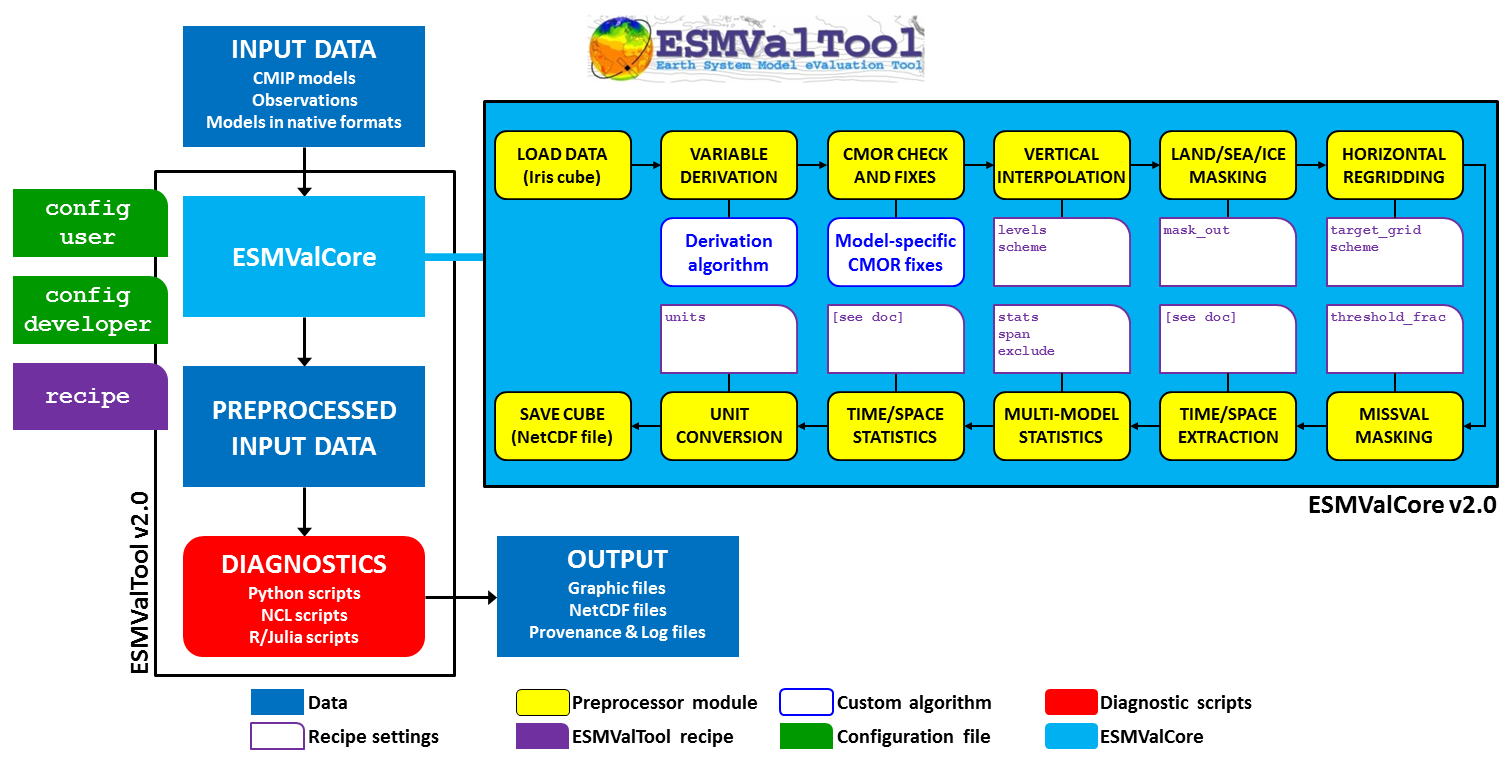
Fig. 1 Schematic of the system architecture.#
For a generalized run scenario, the tool will perform the following ordered procedures.
Data finding#
read the data requirements from the datasets section of the recipe and assemble the data request to locate the data;
find the data using the specified root paths and DRS types in the configuration file (note the flexibility allowed by the data finder);
Data selection#
data selection is performed using the parameters specified in the datasets section (including e.g. type of experiment, type of ensemble, time boundaries etc); data will be retrieved and selected for each variable that is specified in the diagnostics section of the recipe;
Data fixing#
the ESMValTool requires data to be in CMOR format; since errors in the data formatting are not uncommon, the ESMValTool performs checks against the CMOR library and fixes small irregularities (note that the degree of leniency is not very high).
Variable derivation#
variable derivation (in the case of non CMOR-standard variables, most likely associated with observational datasets) is performed automatically before running the preprocessor;
if the variable definitions are already in the database then the user will just have to specify the variable to be derived in the diagnostics section (as any other standard variable, just setting
derive: true).
Run the preprocessor#
if any preprocessor section is specified in the recipe file, then data will be loaded in memory as iris cubes and passed through the preprocessing steps required by the user and specified in the preprocessor section, using the specific preprocessing step parameters provided by the user as keys (for the parameter name) and values (for the parameter value); the preprocessing order is very important since a number of steps depend on prior execution of other steps (e.g. multimodel statistics can not be computed unless all models are on a common grid, hence a prior regridding on a common grid is necessary); the preprocessor steps order can be set by the user as custom or the default order can be used;
once preprocessing has finished, the tool writes the data output to disk as netCDF files so that the diagnostics can pick it up and use it; the user will also be provided with a metadata file containing a summary of the preprocessing and pointers to its output. Note that writing data to disk between the preprocessing and the diagnostic phase is required to ensure multi-language support for the latter.
Run the diagnostics#
the last and most important phase can now be run: using output files from the preprocessor, the diagnostic scripts are executed using the provided diagnostics parameters.

