Shapeselect#
Overview#
Impact modelers are often interested in data for irregular regions best defined by a shapefile. With the shapefile selector tool, the user can extract time series or CII data for a user defined region. The region is defined by a user provided shapefile that includes one or several polygons. For each polygon, a new timeseries, or CII, is produced with only one time series per polygon. The spatial information is reduced to a representative point for the polygon (‘representative’) or as an average of all grid points within the polygon boundaries (‘mean_inside’). If there are no grid points strictly inside the polygon, the ‘mean_inside’ method defaults to ‘representative’ for that polygon. An option for displaying the grid points together with the shapefile polygon allows the user to assess which method is most optimal. In case interpolation to a high input grid is necessary, this can be provided in a pre-processing stage. Outputs are in the form of a NetCDF file, or as ascii code in csv format.
Available recipes and diagnostics#
Recipes are stored in recipes/
recipe_shapeselect.yml
Diagnostics are stored in diag_scripts/shapeselect/
diag_shapeselect.py: calculate the average of grid points inside the user provided shapefile and returns the result as a NetCDF or Excel sheet.
User settings in recipe#
Script diag_shapeselect.py
Required settings (scripts)
shapefile: path to the user provided shapefile. A relative path is relative to the auxiliary_data_dir as configured in config-user.yml.
weighting_method: the preferred weighting method ‘mean_inside’ - mean of all grid points inside polygon; ‘representative’ - one point inside or close to the polygon is used to represent the complete area.
write_xlsx: true or false to write output as Excel sheet or not.
write_netcdf: true or false to write output as NetCDF or not.
Variables#
pr,tas (daily)
Example plots#
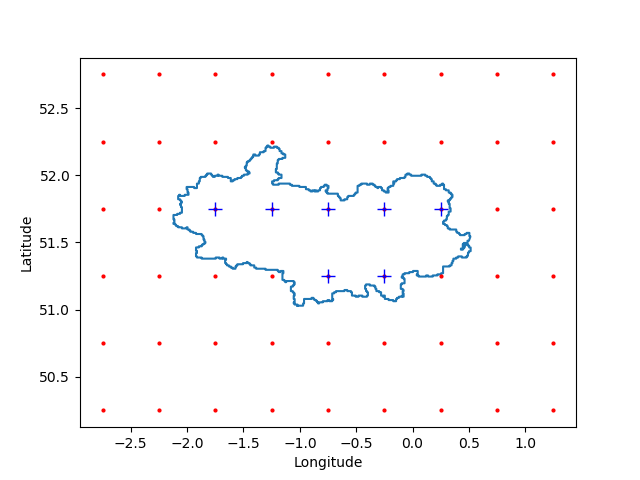
Fig. 244 Example of the selection of model grid points falling within (blue pluses) and without (red dots) a provided shapefile (blue contour).#
