ClimWIP: independence & performance weighting¶
Overview¶
This recipe calculates weights based on combined performance and independence metrics. These weights can be used in subsequent diagnostics. Reference implementation based on https://github.com/lukasbrunner/ClimWIP
Available recipes and diagnostics¶
Recipes are stored in esmvaltool/recipes/
recipe_climwip_test_basic.yml
recipe_climwip_test_performance_sigma.yml
(A recipe adding a ‘real’ case will be added in due time.)
Diagnostics are stored in esmvaltool/diag_scripts/weighting/climwip/
main.py: Compute weights for each input dataset
calibrate_sigmas.py: Compute the sigma values on the fly
core_functions.py: A collection of core functions used by the scripts
io_functions.py: A collection of input/output functions used by the scripts
Plot scripts are stored in esmvaltool/diag_scripts/weighting/
weighted_temperature_graph.py: Show the difference between weighted and non-weighted temperature anomalies as time series.
weighted_temperature_map.py: Show the difference between weighted and non-weighted temperature anomalies as map.
plot_utilities.py: A collection of functions use by the plot scripts.
User settings in recipe¶
Script
main.py
- Required settings for script
performance_sigmaxorcalibrate_performance_sigma: Ifperformance_contributionsis given exactly one of the two has to be given. Otherwise they can be skipped or not set.
performance_sigma: float setting the shape parameter for the performance weights calculation (determined offline).
calibrate_performance_sigma: dictionary setting the performance sigma calibration. Has to contain at least the key-value pair specifyingtarget:variable_group. Optional parameters for adjusting the calibration are not yet implemented. WARNING: It is highly recommended to visually inspect the graphical output of the calibration to check if everything worked as intended. In case the calibration fails, the best performance sigma will still be indicated in the figure (see example Fig. 8 below) but not automatically picked - the user can decide to use it anyway by setting it in the recipe (not recommenced).
independence_sigma: float setting the shape parameter for the independence weights calculation (determined offline). Can be skipped or not set ifindependence_contributionsis skipped or not set.
performance_contributions: dictionary where the keys represent the variable groups to be included in the performance calculation. The values give the relative contribution of each group, with 0 being equivalent to not including the group. Can be skipped or not set then weights will be based purely on model independence (this is mutually exclusive withindependence_contributionsbeing skipped or not set).
independence_contributions: dictionary where the keys represent the variable groups to be included in the independence calculation. The values give the relative contribution of each group, with 0 being equivalent to not including the group. Can be skipped or not set then weights will be based purely on model performance (this is mutually exclusive withperformance_contributionsbeing skipped or not set).
combine_ensemble_members: set to true if ensemble members of the same model should be combined during the processing (leads to identical weights for all ensemble members of the same model). Recommended if running with many (>10) ensemble members per model.
obs_data: list of project names to specify which are the the observational data. The rest is assumed to be model data.- Required settings for variables
This script takes multiple variables as input as long as they’re available for all models
start_year: provide the period for which to compute performance and independence.
end_year: provide the period for which to compute performance and independence.
mip: typically Amon
preprocessor: e.g. climwip_summer_mean
additional_datasets: provide a list of model data for performance calculation.- Required settings for preprocessor
Different combinations of preprocessor functions can be used, but the end result should always be aggregated over the time dimension, i.e. the input for the diagnostic script should be 2d (lat/lon).
- Optional settings for preprocessor
extract_regionorextract_shapecan be used to crop the input data.
extract_seasoncan be used to focus on a single season.different climate statistics can be used to calculate mean or (detrended) std_dev.
Script
weighted_temperature_graph.py
- Required settings for script
ancestors: must include weights from previous diagnostic
weights: the filename of the weights: ‘weights.nc’- Required settings for variables
This script only takes temperature (tas) as input
start_year: provide the period for which to plot a temperature change graph.
end_year: provide the period for which to plot a temperature change graph.
mip: typically Amon
preprocessor: temperature_anomalies- Required settings for preprocessor
Different combinations of preprocessor functions can be used, but the end result should always be aggregated over the latitude and longitude dimensions, i.e. the input for the diagnostic script should be 1d (time).
- Optional settings for preprocessor
Can be a global mean or focus on a point, region or shape
Anomalies can be calculated with respect to a custom reference period
Monthly, annual or seasonal average/extraction can be used
Script
weighted_temperature_map.py- Required settings for script
ancestors: must include weights from previous diagnosticweights: the filename of the weights: ‘weights_combined.nc’
- Optional settings for script
model_aggregation: how to aggregate the models: mean (default), median, integer between 0 and 100 given a percentilexticks: positions to draw xticks atyticks: positions to draw yticks at
- Required settings for variables
This script takes temperature (tas) as input
start_year: provide the period for which to plot a temperature change graph.end_year: provide the period for which to plot a temperature change graph.mip: typically Amonpreprocessor: temperature_anomalies
- Optional settings for variables
A second variable is optional: temperature reference (tas_reference). If given, maps of temperature change to the reference are drawn, otherwise absolute temperature are drawn.
tas_reference takes the same fields as tas
Variables¶
pr (atmos, monthly mean, longitude latitude time)
tas (atmos, monthly mean, longitude latitude time)
psl (atmos, monthly mean, longitude latitude time)
more variables can be added if available for all datasets.
Observations and reformat scripts¶
Observation data is defined in a separate section in the recipe and may include multiple datasets.
References¶
Example plots¶
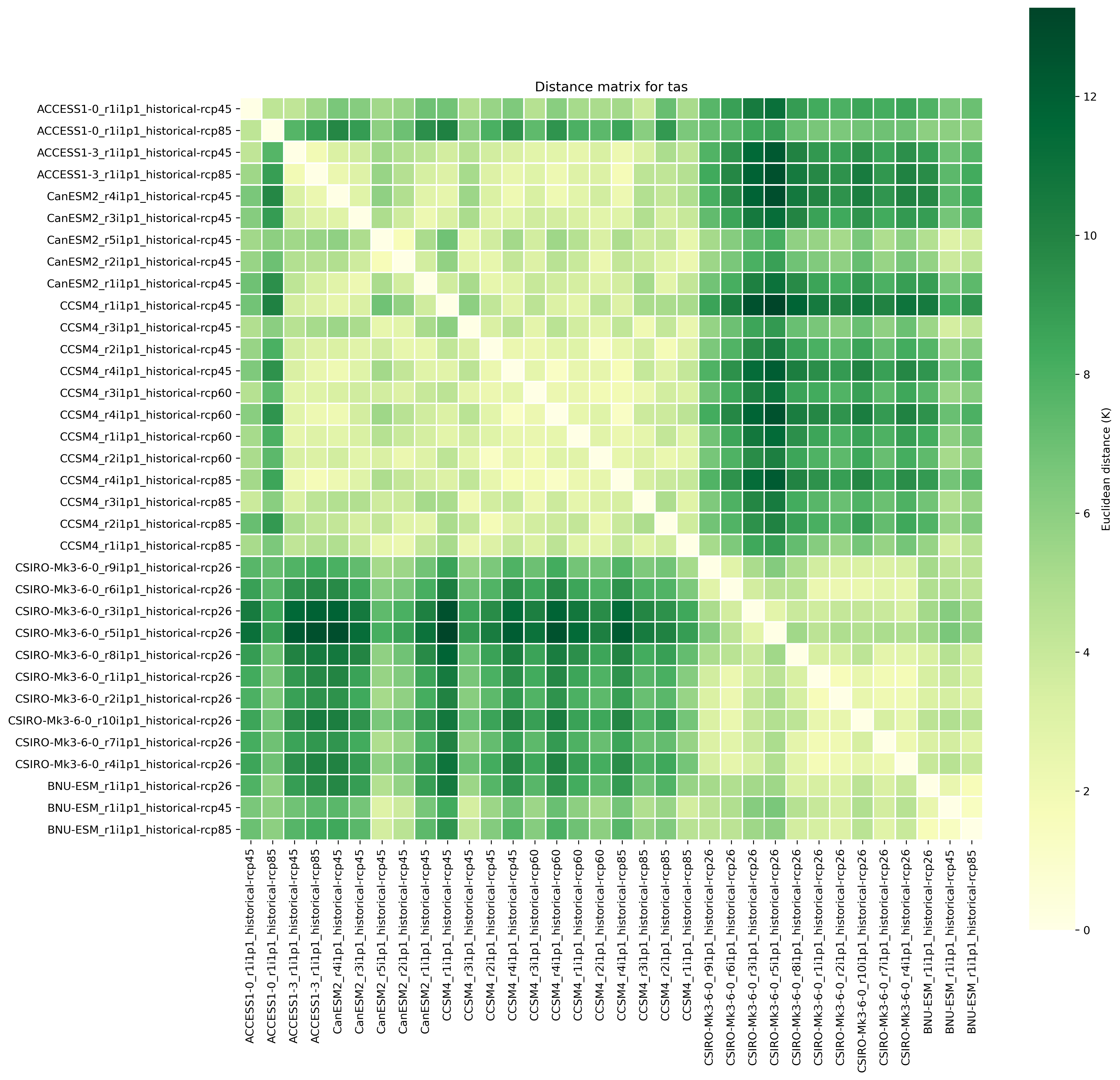
Fig. 4 Distance matrix for temperature, providing the independence metric.¶
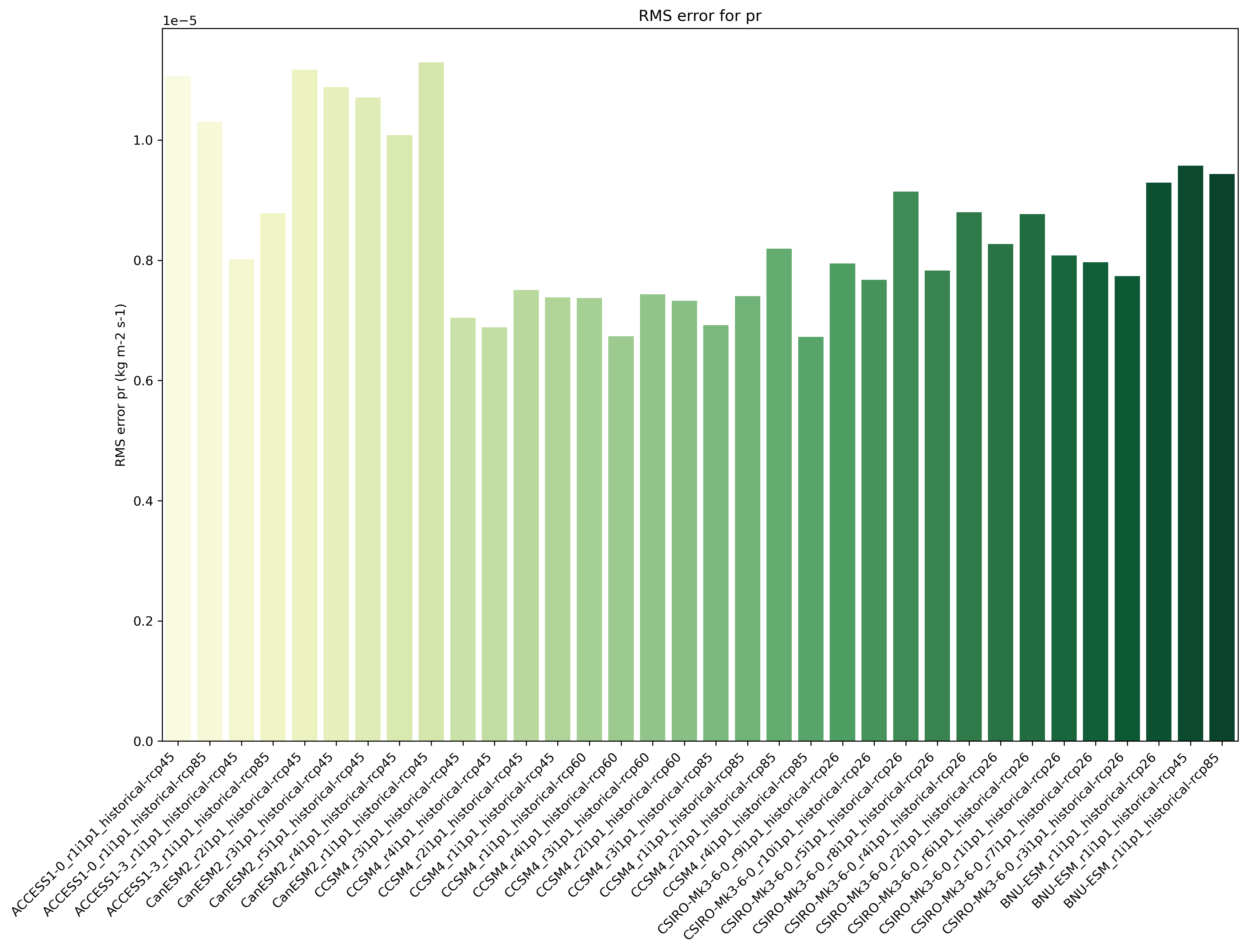
Fig. 5 Distance of preciptation relative to observations, providing the performance metric.¶
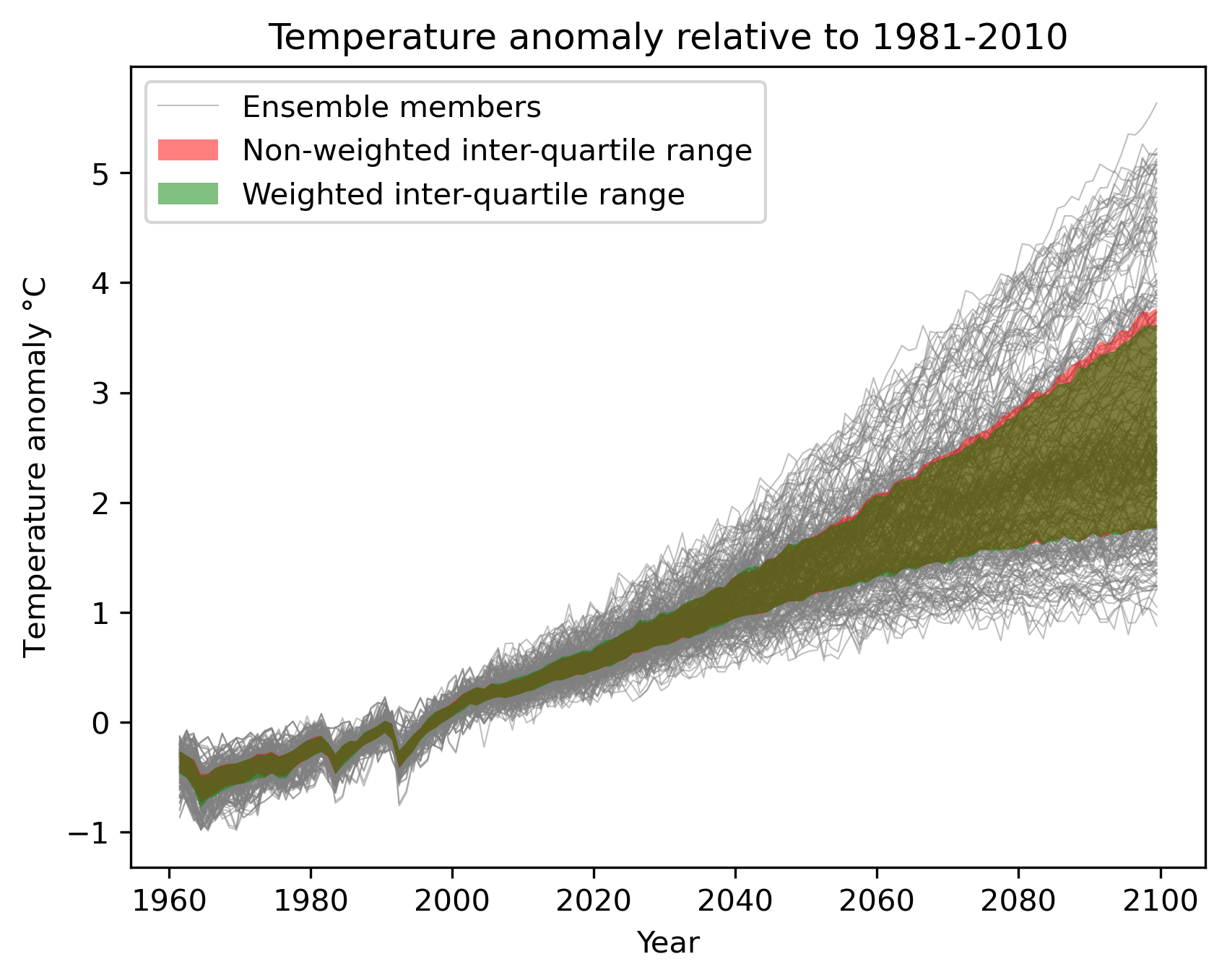
Fig. 7 Interquartile range of temperature anomalies relative to 1981-2010, weighted versus non-weighted.¶
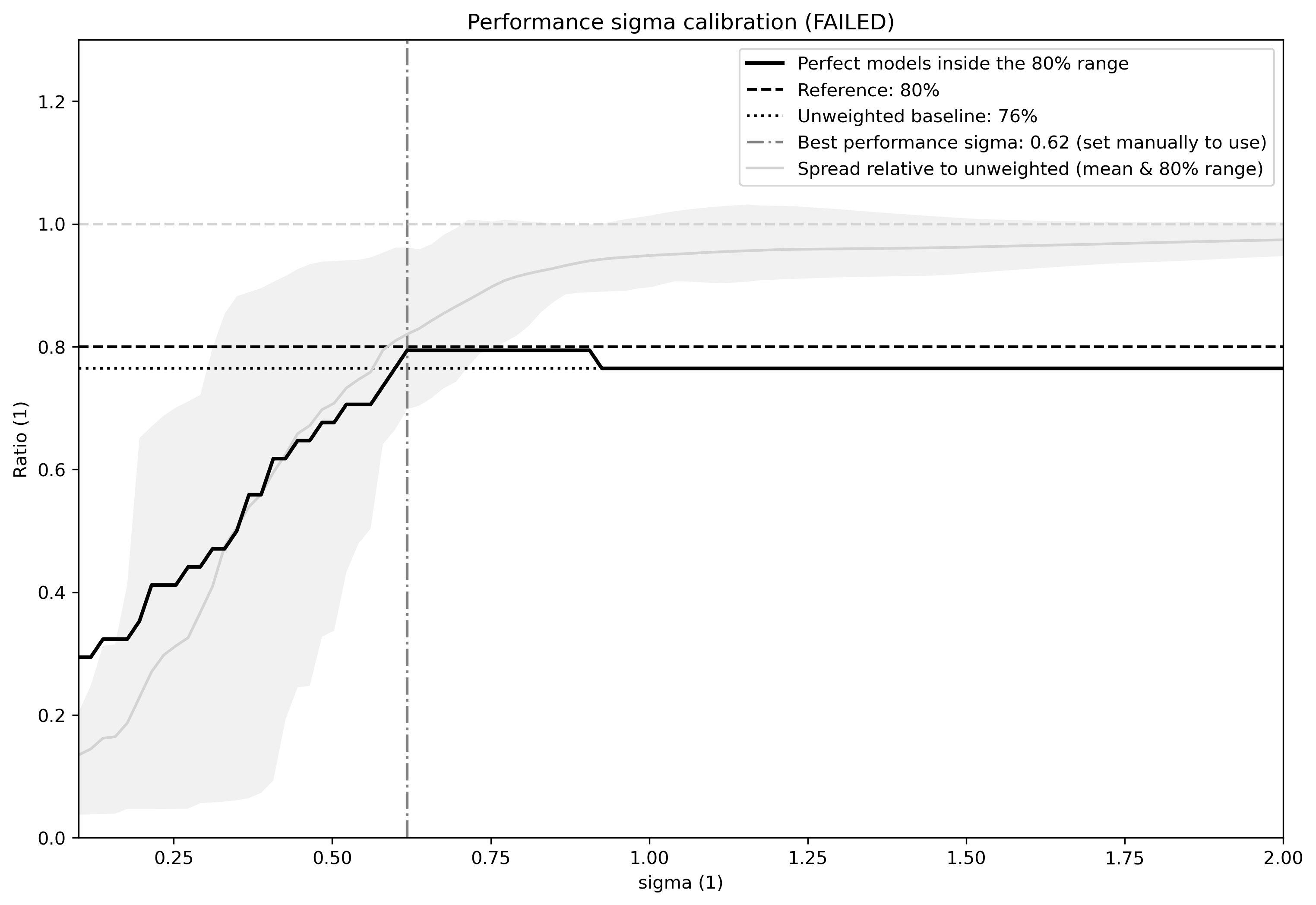
Fig. 8 Performance sigma calibration: The thick black line gives the reliability (c.f., weather forecast verification) which should reach at least 80%. The thick grey line gives the mean change in spread between the unweighted and weighted 80% ranges as an indication of the weighting strength (if it reaches 1, the weighting has no effect on uncertainty). The smallest sigma (i.e., strongest weighting) which is not overconfident (reliability >= 80%) is selected. If the test fails (like in this example) the smallest sigma which comes closest to 80% will be indicated in the legend (but NOT automatically selected).¶
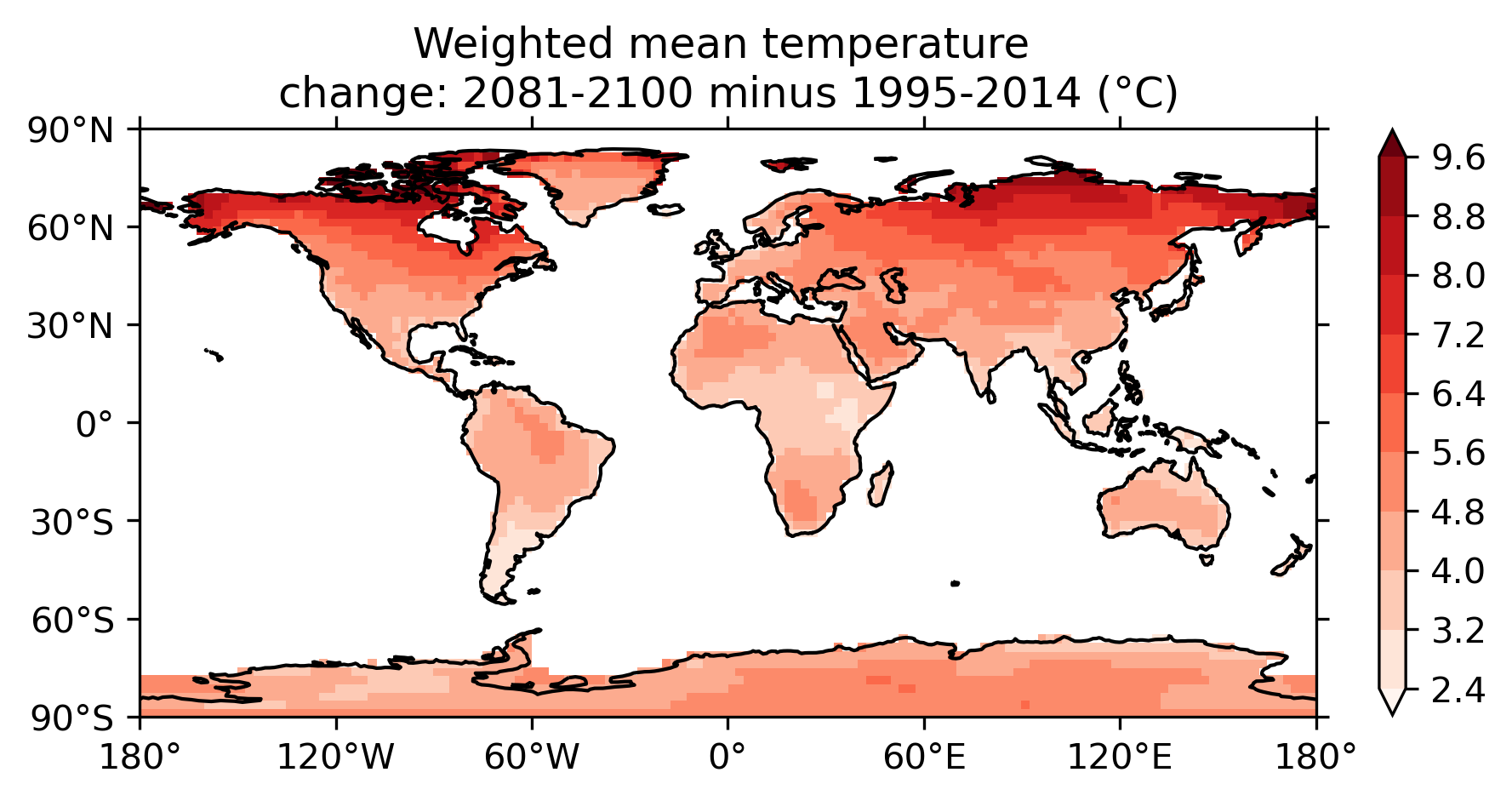
Fig. 9 Map of weighted mean temperature change 2081-2100 relative to 1995-2014¶